Spectrum of methods
To assess the influence of metals and metal compounds, nanomaterials and bioactive food compounds on genomic stability in cell culture a broad range of different methods was established in our laboratory. They include:
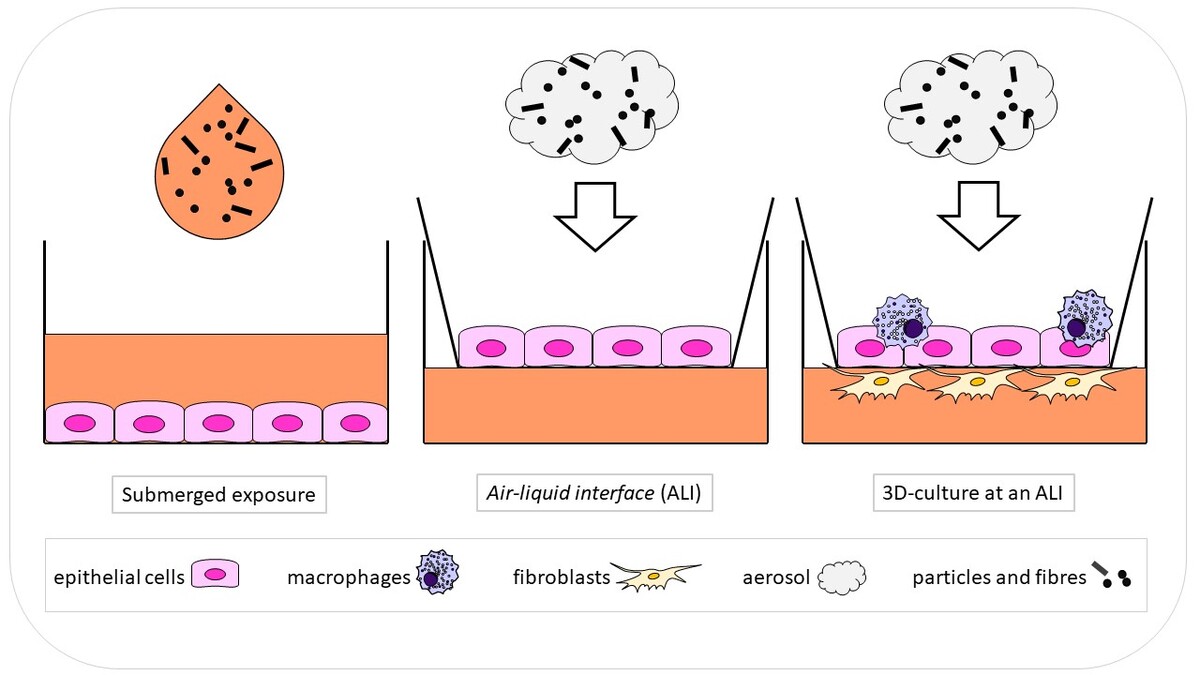
- the application of regular, co- and 3D-cell culture systems
- determination of viability and cytotoxicity,
- examination of apoptosis and cell cycle control (e.g. with flow cytometry),
- measuring concentrations of selected metals with atom absorption spectrometry (AAS) and colorimetric methods,
- quantitative and qualitative protein determination with Western Blot, immunofluorescence and ELISA,
- the quantification of DNA damage and repair induction as well as mutagenicity
- oxidative DNA damage, benzo[a]pyrene-induced DNA damage, UVC-induced DNA damage, DNA single and double strand breaks, e.g. by alkaline unwinding, micronuclei induction, HPLC with fluorescence detection,
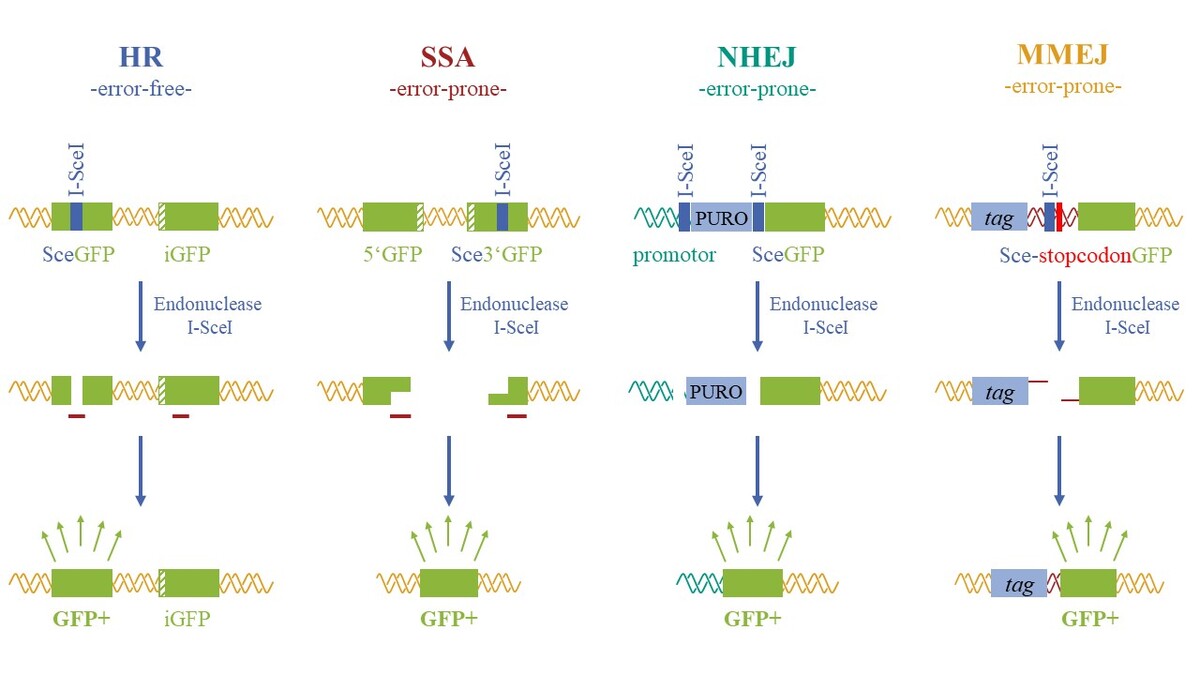
- examination of different DNA repair pathways (e.g. wiith reporter test systems, repair-deficient cell lines, accumulation of DNA repair proteins),
- examination of PARP activity,
- determination of mutagenicity by PIG-A assay,
- the observation of selected cellular processes with live cell imaging (in a time-dependent manner),
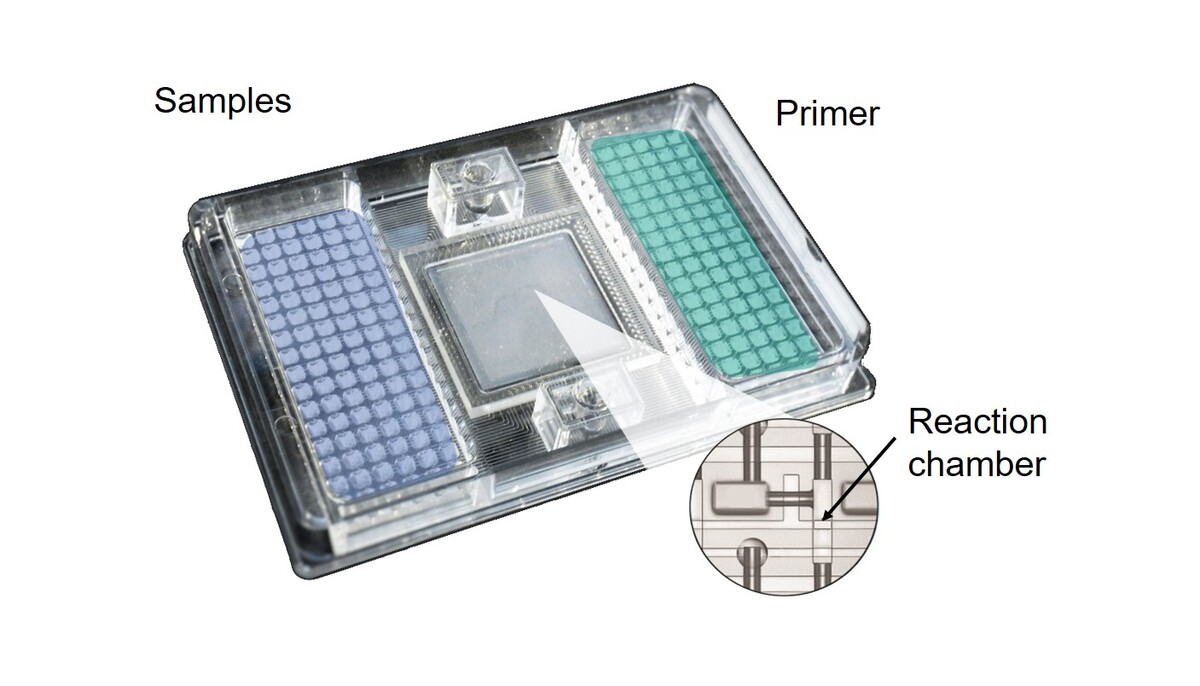
- the analysis of gene expression profiles with real-time RT-PCR and multiplex-PCR
- influence on genomic stability (oxidative stress, general stress response, DNA damage response, cell cycle control, metal homeostasis, inflammatory and fibrotic responses),
- the examination of particular and fibrous nanomaterials
- characterization of nanomaterials,
- air-liquid interface exposure,
- comparison of soluble compounds with nano- and microparticles as well as fibrous materials with regard to cytotoxicity, DNA damage, gene expression profiling and other endpoints.